|
gatb.core-API-0.0.0
|
|
gatb.core-API-0.0.0
|
As its name suggests, the gatb::core package holds all the fundamental packages needed for writting assembly algorithms.
It holds some generic services, like:
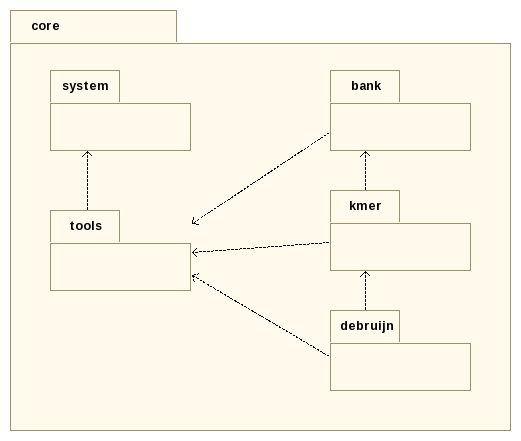
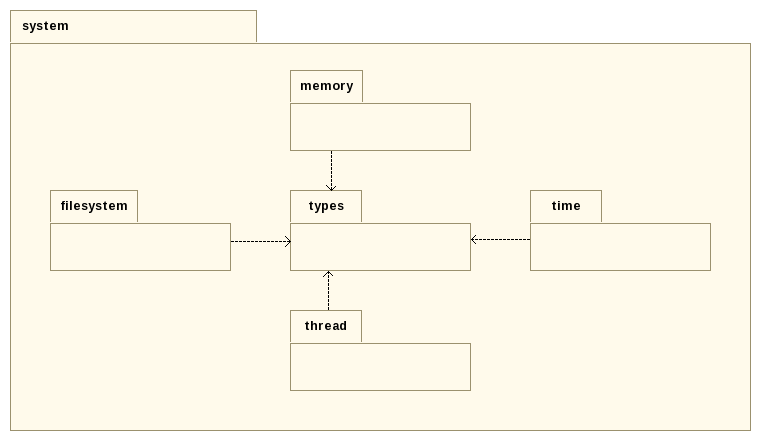
The gatb::core::system package holds operations related to operating system.
Making such an abstraction is interesting when considering the following points:
The idea is so to list all necessary operating system operations that are likely to be used, to make several interfaces that gather some operations set and to provide specific implementations for several system (linux, macos, windows).
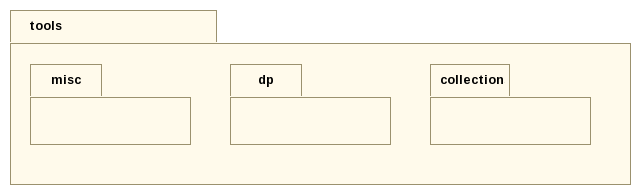
The gatb::core::tools package holds generic operations used throughout the project. They are not specific to genome nor assembly concerns and are defined here in a separate package.
Roughly speaking, we may find the following operations:
The gatb::core::bank package holds operations related to genomic databases management.
In particular, we define a gatb::core::bank::IBank interface that is an iterable over gatb::core::bank::Sequence instances. This interface has a lot of implementations, including FASTA format.
The gatb::core::kmer package holds operations related to kmer management.
A typical case is to create a IBank instance and then an iterator on it; such an iterator can feed the kmer model for iterating all the kmer from the sequences.
The gatb::core::debruijn package holds operations related to De Bruijn graph.
The main API is the gatb::core::debruijn::impl::Graph class. It holds several features:
The implementation of the class relies on a probabilistic data structure named Bloom filter. Its usage allows to have a very low memory footprint, even for big graphs.